购物车
 您的购物车当前为空
您的购物车当前为空
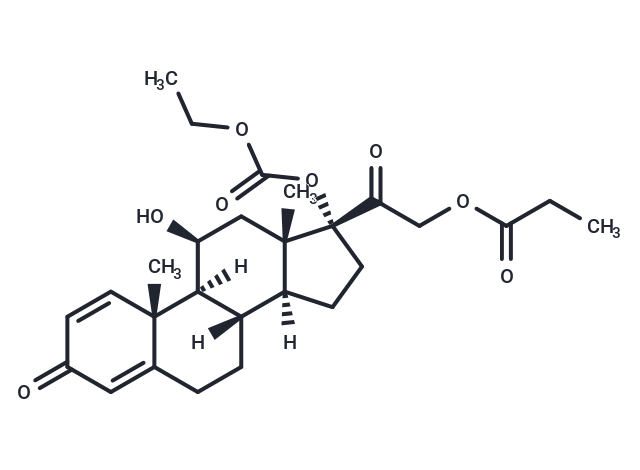
Prednicarbate (Hoe 777) 是一种外用皮质类固醇。Prednicarbate 可用于炎症性皮肤病的研究,如特应性皮炎。
别名 泼尼卡酯, UNII-V901LV1K7D, Hoe-777, Hoe 777, Dermatop E emollient
Prednicarbate (Hoe 777) 是一种外用皮质类固醇。Prednicarbate 可用于炎症性皮肤病的研究,如特应性皮炎。
| 规格 | 价格 | 库存 | 数量 |
|---|---|---|---|
| 1 mg | ¥ 248 | 现货 | |
| 5 mg | ¥ 558 | 现货 | |
| 10 mg | ¥ 845 | 现货 | |
| 25 mg | ¥ 1,180 | 现货 | |
| 50 mg | ¥ 1,650 | 现货 | |
| 100 mg | ¥ 2,480 | 现货 | |
| 1 mL x 10 mM (in DMSO) | ¥ 588 | 现货 |
TargetMol的所有产品仅用作科学研究或药证申报,不能被用于人体,我们不向个人提供产品和服务。请您遵守承诺用途,不得违反法律法规规定用于任何其他用途。

| 产品描述 | Prednicarbate (Hoe 777) is a Corticosteroid Hormone Receptor Agonist. Prednicarbate has antipruritic, anti-inflammatory, and vasoconstrictive properties. It also decreases the number of circulating lymphocytes by inhibiting the production of vasodilators such as nitric oxide and prostacyclin. |
| 别名 | 泼尼卡酯, UNII-V901LV1K7D, Hoe-777, Hoe 777, Dermatop E emollient |
| 分子量 | 488.57 |
| 分子式 | C27H36O8 |
| CAS No. | 73771-04-7 |
| Smiles | C(COC(CC)=O)(=O)[C@]1(OC(OCC)=O)[C@]2(C)[C@@](CC1)([C@]3([C@]([C@@H](O)C2)([C@]4(C)C(CC3)=CC(=O)C=C4)[H])[H])[H] |
| 密度 | 1.25 g/cm3 |
| 存储 | Powder: -20°C for 3 years | In solvent: -80°C for 1 year Shipping with blue ice/Shipping at ambient temperature. 实际储存温度请以COA为准 | |||||||||||||||||||||||||
| 溶解度信息 | H2O: Insoluble DMSO: 10 mg/mL (20.47 mM), Sonication is recommended. | |||||||||||||||||||||||||
溶液配制表 | ||||||||||||||||||||||||||
DMSO
该溶液配制表仅适用于固体产品。对于液体产品,请根据标明的浓度或密度计算稀释方案。 | ||||||||||||||||||||||||||
对于不同动物的给药剂量换算,您也可以参考 更多